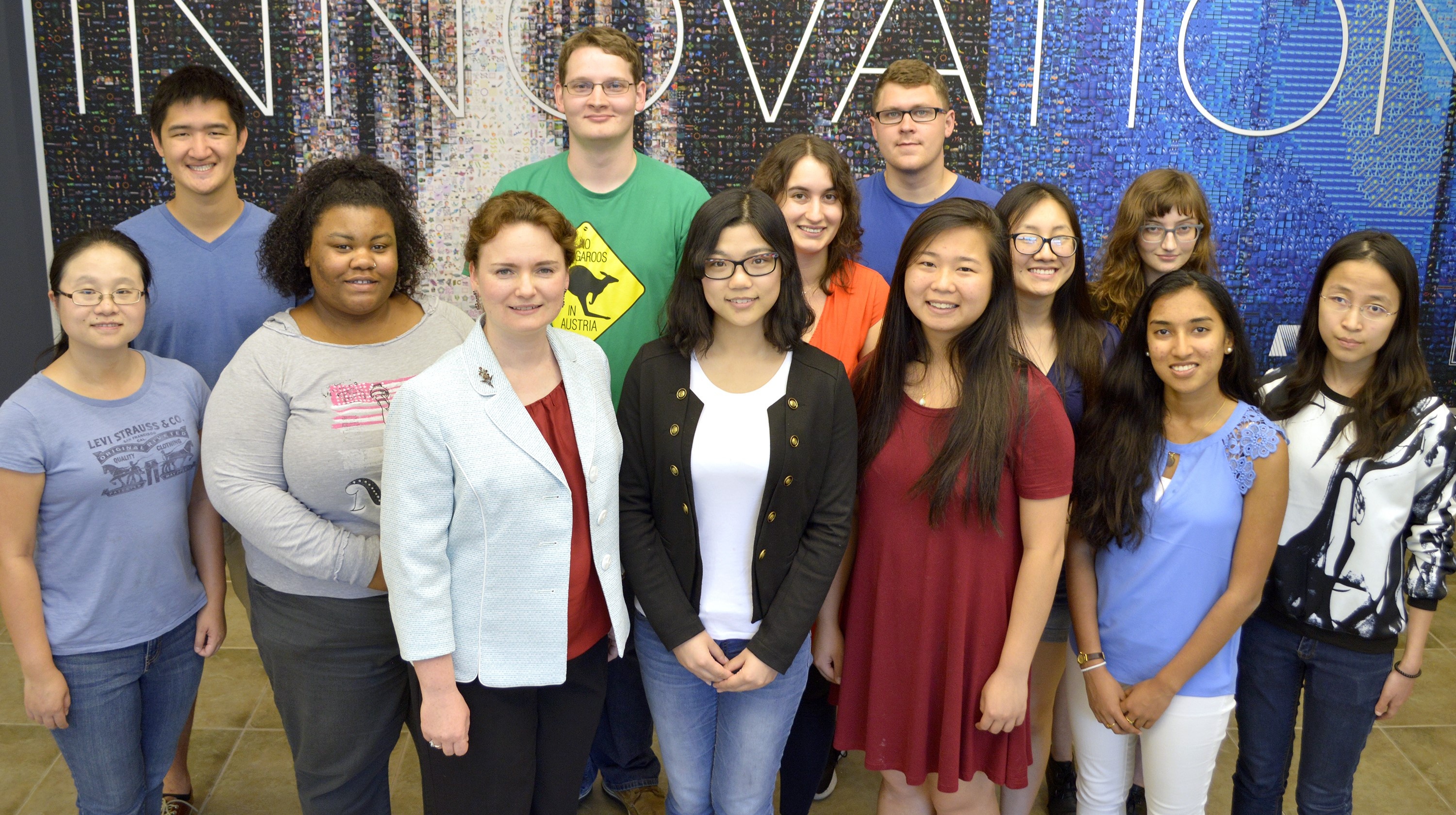
This group is a host for research into the use of high performance computing (HPC) for primary genomics analyses, such as alignment, variant calling, genome assembly, and RNASeq. By its nature, this research is highly collaborative. Every member of our team is affiliated with multiple departments or campus initiatives. The student participants in this group serve as a bond between the campus faculty using computational genomics analyses in their research, and the NCSA experts in HPC, storage, networking, databases, etc. Together we enable the use of advanced computing infrastructure in computational genomics. Explore this page to find out who is involved, how we are connected, and what projects are currently ongoing.
NCSA Press: Crossing over, branching out: Meet the NCSA Genomics team
Technical Program Manager, National Center for Supercomputing Applications
Research Assistant Professor, Institute of Genomic Biology
217-300-0568
NCSA Genomics, September 2017. Credit: Steve Deunsing
Not pictured: Matt Weber, Ram Venkatakrishnan
Current Projects | ||
B.S. Molecular and Cellular Biology (2016) M.S. Bioinformatics (2018) Department of Crop Sciences, UIUC CompGen fellow advised by Dr. Matthew Hudson | Mutation profiles of cancerMr. Weber is developing machine learning methods to effectively stratify cancers based on the Paper: Simulating Next-Generation Sequencing Datasets from Empirical Mutation and Sequencing Models Poster: Statistical models to capture mutational properties for NextGen Sequencing Data | |
B.S. Crop Science (2016) Plant Biotechnology, Molecular Biology M.S. Bioinformatics (2018) Department of Crop Sciences, UIUC Graduate Fellow in the College of ACES advised by Dr. Matthew Hudson | Genomic variant calling by assemblyMr. K is focusing on a method to detect genomic variants by assembly. He is employing the software Cortex-var, which constructs de-novo genome assembly on multiple Mr. K is also working with Tiffany on the genomic analysis of HLHS for the Mayo Grand Challenge. Poster: Variant Calling by Assembly Poster: Reference-guided variant calling for non-repetitive sequences in Glycine Max | |
Tiffany Li B.S. Integrative Biology (2018) minor in Computer Science | Benchmarking performance and accuracy of genomic variant calling softwareTiffany collaborates to document our efforts in benchmarking variant calling on HPC We have also tested a number of alternative software, such as Isaac, Genalice, and Sentieon, Validation and benchmarking on ParFu - a parallel file packaging utilityTiffany is also involved in testing and benchmarking of ParFu, an MPI tool for creating or extracting | |
B.S. Biochemistry (2014) M.S. Bioinformatics (2016) Bioinformatics specialist and research programmer at NCSA | NCSA IndustryJacob supports biomedical partners in the NCSA Industry program. Jacob provides a complementary mix of expertise in HPC and bioinformatics data analysis to enable Jacob and Azza Ahmed (Ph. D. candidate, University of Khartoum) are exploring and evaluating the | |
Aishwarya Raj B.S. Biochemistry (2019) minor in Bioinformatics | Evolution of molecular networks and persistence of organismsConstruct and compare gene, metabolic and signaling networks from organisms across the tree of life. The goal of the project is to provide support for the general framework of persistence strategies. It postulates that persistence is achieved by biological systems via a tradeoff of traits that serve either Poster: Persistence Strategies in Biomolecular Network Architecture | |
Ellen Nie B.S. Computer Science (2018) | Big data network transfers for genomicsEllen is benchmarking the network transfers of genomic data across multiple sites. Poster: Benchmarking and Optimization of Long Distance Big Data Transfers Validation of Sentieon - the fast alternative to GATKEllen is also collaborating with OICR to validate the speed and accuracy of the new software Convert Java-based GWAS code for SparkIn a project described below (Accurate and scalable GWAS algorithms) we are improving performance of We would like to deploy this Java code on Spark, to see if the necessary performance gains could be obtained. A successful student applicant will use Java Spark API to adapt the current code for a Spark platform that Poster: Scaling the Computation of Epistatic Interactions in GWAS Data | |
Weihao Ge B.S. Physics (2008) M.S. Physics (2011) Ph.D. Biophysics (2018) advised by Dr. Eric Jacobsson | Search Space ReductionWeihao is evaluating statistical methods for search space reduction in the analysis of GWAS data for genomic variant | |
Ramshankar Venkatakrishnan B.S. Electronics & Communications (2012) M.S. Electrical & Computer Engineering (2015) | Mayo Grand Challenge: evaluating and streamlining genomics workflowsRamshankar and Cynthia are working on computational improvements for the Mayo Grand Challenge, a genomics research Ram will also contribute his hardware expertise to the project, evaluating system architecture options to complement the team’s | |
Cynthia Liu B.S. Bioengineering (2019) minor in Computer Science | ||
Sijia Huo B.S. Mathematics & Computer Science (2018) second major in Statistics third major in Economics | Parallelization of RSijia is working with NCSA Faculty Fellow Dr. Zeynep Madak-Erdogan to introduce parallel R code into her research. Dr. Madak-Erdogan is exploring racial disparities in breast cancer occurrence through the lens of diet and nutrition. | |
Katherine Kendig B.A. Anthropology (2012) M.F.A. Creative Writing (2017) | Research CoordinationKatherine assists the team with scheduling, manuscript editing and preparation, documentation, and project communications. She also contributes to NCSA’s Public Affairs team. | |
Former Group Members | ||
Ryan Chui B.S. Biochemistry (2016) M.S. Bioinformatics (2017) | NCSA IndustryRyan performed software installation, benchmarking, and development for a variety of industry partners. To investigate how the training time for deep neural networks (DNN’s) can be affected, Ryan worked On Github: EpiQuant: Hadoop, C, Tensorflow - epistasis software prototypes MLCC - multi-label cancer classification q2b - binary representation of nucleotides ptgz - parallel tar gzip Usage Analyzer - log analyzer for HPC schedulers | |
Jennie Zermeno B.S. Integrative Biology (2017) | Benchmarking performance and accuracy of genomic variant calling softwareJennie collaborated to document our efforts in benchmarking variant calling on HPC Bioinformatics in the CloudJennie is investigating the issues of portability, reproducibility and scaling of bioinformatics workflows Students Capitalize on Computational Genomics Research Using AWS | |
Angela Chen M.S. Statistics (2017) Department of Statistics, UIUC CompGen fellow advised by Dr. Alexander Lipka | Accurate and scalable GWAS algorithmsAngela and Khory collaborated to improve the scalability and parallelization of the statistical software Angela wrote a manuscript to demonstrate that her new stepwise epistatic model selection Khory provided the expertise in computer science to convert this Java code into C++ and parallelize | |
Khory Wagner advised by Dr. Vologymyr Kindratenko | ||
Nainika Roy B.S. Molecular and Cellular Biology (2017) minor in Informatics and Chemistry SPIN fellow | Data formats and data structures in computational genomics | |
Junyu Li B.S. Molecular and Cellular Biology (2017) minor in Computer Science SPIN fellow | Genomic variant calling by assemblyJunyu worked with Mr. K in an interdisciplinary team, providing the expertise in math and computer Poster: Reference-guided variant calling for novel non-repetitive sequences in Glycine max | |
Noah Flynn B.S. Bioengineering, Mathematics (2017) minor in computer science SPIN fellow | Evolution of molecular networks and persistence of organisms | |
Other Collaborations
Dr. Matthew Hudson Bioinformatics Crop Science | HPCBio, Carver Biotechnology Center | |
Dan Wickland Ph.D. Informatics (2019) | ||
Computer Science | NCSA Scientific Software and ApplicationsPortable variant calling workflow in Swift
| |
Azza Ahmed Computer Science advised by Dr. Faisal Fadlelmola | ||
Dr. Zeynep Madak-Erdogan Food Science & Human Nutrition | Madak-Erdogan LabSystems Biology of Estrogen Signaling
| |
Brandi Smith Ph.D. Food Science and Human Nutrition (2021) | ||
H3Africa Consortium
| ||
Morgan Taschuk Bioinformatics | OICR
| |
Paul Hatton HPC / Visualisation
| University of Birmingham |