Motivation
A couple basic tools are included with most linux distributions providing code coverage analysis and function profiling: gcov and gprof. Using just these 2 tools, it's easy to gain understanding of how your code is running. With the linuxtools Eclipse incubation project ( http://www.eclipse.org/linuxtools/ , now bundled with PTP ) , gprof and gcov are fully integrated into the cdt editor perspective. Let's see how the linuxtools project adds value and puts a fresh look on these classic linux code analysis tools.
Demo : Shallow water weather model
The Shallow demo project from the eclipse tutorial cvs can showcase gprof and gcov with a few simple adjustments to the Makefile and MPI runtime. Here are the details and some discussion of what can be done with the profiling and coverage output.
First, the source code is from the demo project and the checkout is described in Module 3 of the PTP SC11 Tutorial.
Once checked out, fix the bug in the for() loop of main.c and build the project (Copy Makefile.mk to Makefile and adjust as needed). Then setup a run configuration with openmpi on your favorite system. I've chosen my own laptop but you could also setup a synchronized project and use a remote system of your choice. My laptop runs Ubuntu, the GNU compilers, and OpenMPI which makes working with Eclipse and developing test/demo codes a snap.
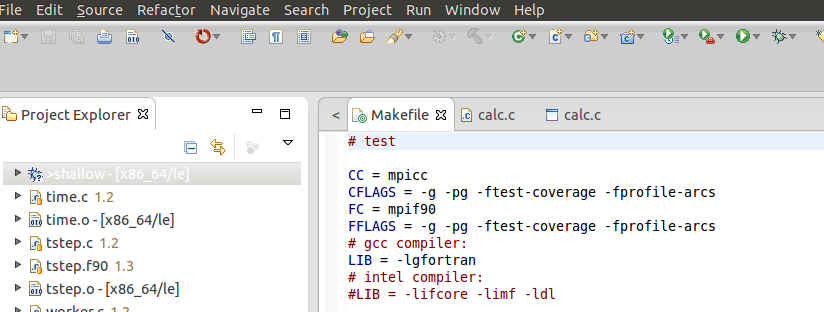
To use gprof and gcov, add the appropriate flags to the Makefile. I'll run a single test job to get both profile and coverage data at once by using -pg for gprof and "-ftest-coverage -fprofile-arcs" for gcov (-g is included so that we have source code line numbering available). Do a project .> refresh and note the additional files created for use by gcov after compiling with "-ftest-coverage -fprofile-arcs".
Setup the MPI run configuration with the Environment variable GMON_OUT_PREFIX defined with a name for your individual MPI rank gmon outputs. By default gmon.out is used but MPI doesn't do that well and you end up with a profile that's missing most of the information, so by using GMON_OUT_PREFIX, each MPI rank adds its process id to its gmon output filename.
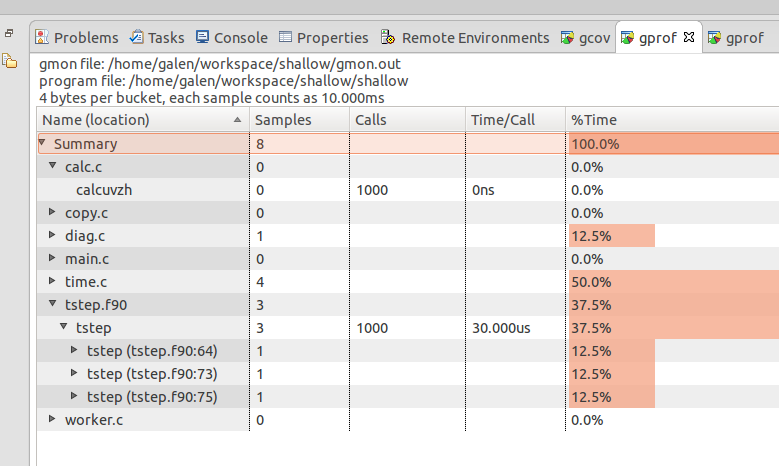
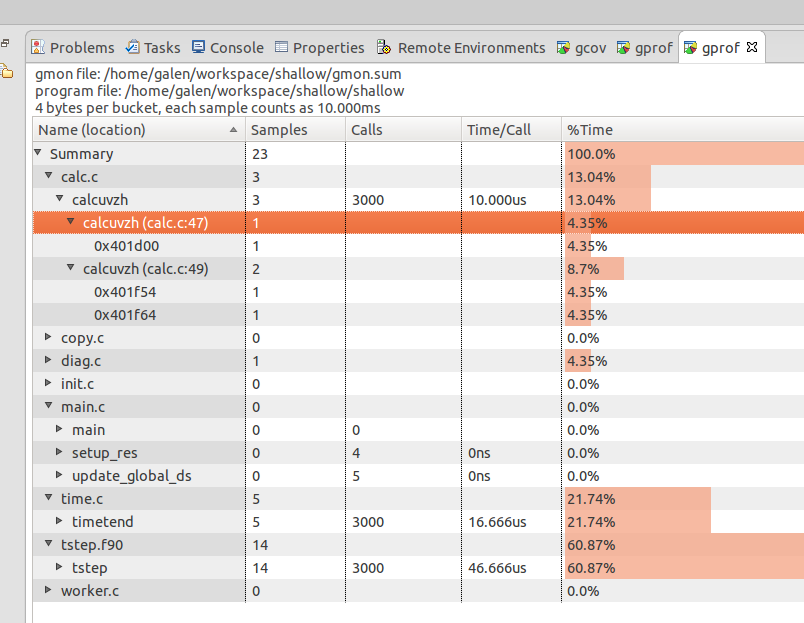
The gmon output files can be combined in a summary with the gprof -s command as shown. It's interesting to compare the summary gmon output to that from one of the ranks (copy a rank's gmon_* to gmon.out to easily view it with Eclipse linuxtools ).
galen@lgheron:~/workspace/shallow$ gprof -s shallow gmon_* galen@lgheron:~/workspace/shallow$ ls calc.c decs.h eclipse.inc init.o time.c worker.c calc.gcda diag.c gmon_shallow.9885 main.c time.gcda worker.gcda calc.gcno diag.gcda gmon_shallow.9886 main.gcda time.gcno worker.gcno calc.o diag.gcno gmon_shallow.9887 main.gcno time.o worker.o copy.c diag.o gmon_shallow.9888 main.o tstep.c copy.gcda dump.c gmon.sum Makefile tstep.f90 copy.gcno dump.gcda init.c Makefile.gem tstep.gcda copy.o dump.gcno init.gcda Makefile.mk tstep.gcno CVS dump.o init.gcno shallow tstep.o galen@lgheron:~/workspace/shallow$
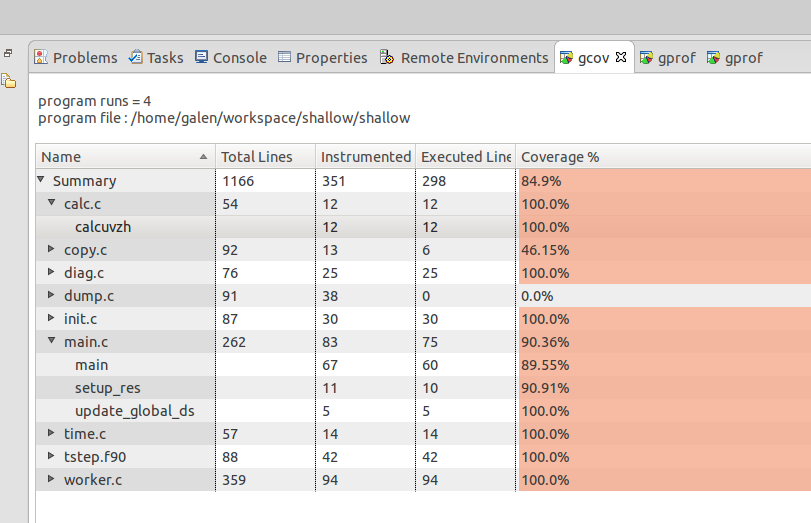
The gcov view is simlar to the gprof view but keep in mind that you're looking at code coverage and not necessarily performance or timing information (though there is a relationship...code not executed is performing quite well ! ). Also note that multiple executions will accumulate values in the gcov output files until they are removed or truncated to zero-length (2nd run to demonstrate this).
Selecting (double click) a source code line from either the gcov or gprof view and you'll see the file and routine highlighted in the cdt c/c++ perspective. Also notice the support for the .f90 file and its routines.
References:
http://wiki.eclipse.org/Linux_Tools_Project/GProf/User_Guide
http://www.ncsa.illinois.edu/UserInfo/Resources/Hardware/SGIAltix/Doc/timing.html
http://archive.ncsa.illinois.edu/lists/perftools/may02/msg00001.html